research
Left-right asymmetry in Drosophila
Analysis of planar cell-shape chirality
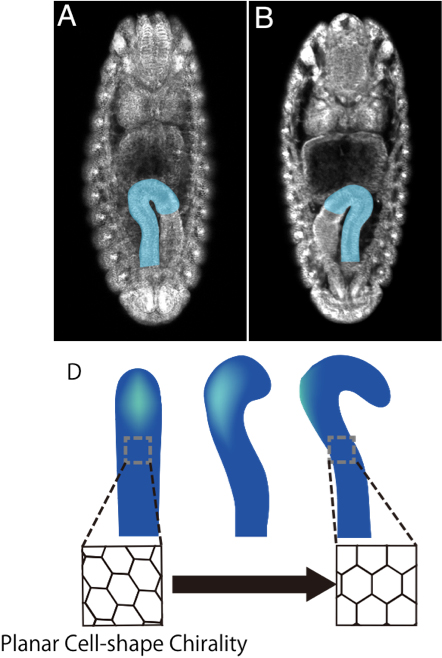
We found that the hindgut epithelial tube rotates 90 degree counter clockwise. During this process, cell-proliferation and apoptosis are not observed, suggesting that cell-rearrangements or cell-shape change play a crucial role. Our imaging analysis of the hindgut epithelial cells revealed that their shape of apical plane shows LR asymmetry.
Their LR asymmetric structure does not overlap with its mirror image. Therefore, we refer to this novel cell behaviour as “planar cell-shape chirality” (PCC). Consistent with this observation, we found that the position of the centrosome also showed LR bias in the hindgut epithelial cells. Importantly, PCC was independently reported in mammalian cultured cells. This suggests that PCC may be evolutionarily conserved.
In Myo31DF homozygote, LR asymmetry of the embryonic hindgut was reversed (become mirror image of the wild-type). In these embryos, PCC was reversed. In DE-Cad homozygote, the laterality of this organ was randomized. We found that PCC was randomized in these embryos. In the epithelial cells of the embryonic hindgut, the distribution of DE-Cad was aligned LR asymmetrically in Myo31DF-dependent manner, which probably contributed to normal PCC formation. Currently, we are studying the molecular mechanisms of the PCC formation.
In Myo31DF homozygote, LR asymmetry of the embryonic hindgut was reversed (become mirror image of the wild-type). In these embryos, PCC was reversed. In DE-Cad homozygote, the laterality of this organ was randomized. We found that PCC was randomized in these embryos. In the epithelial cells of the embryonic hindgut, the distribution of DE-Cad was aligned LR asymmetrically in Myo31DF-dependent manner, which probably contributed to normal PCC formation. Currently, we are studying the molecular mechanisms of the PCC formation.
Identification of genes responsible for the LR asymmetric development
To identify genes responsible for developing the LR asymmetry of the Drosophila embryonic gut, we conducted large-scale genetic screens for mutants on the 2nd and 3rd chromosomes, which were induced by EMS or P-element insertions. We have screened about 8,000 mutant lines in total. More recently, we also performed a maternal screen for mutations on the left-arm of the second chromosome.
In total, about 20 genes that affect the LR asymmetric development of the embryonic gut were identified. The general morphology and tissue-specification of the gut was not affected in these mutants.
Among these mutation, Myosin31DF (Myo31DF) (also called Myosin ID) and Drosophila E-Cadherin (DE-Cad) affect the LR symmetry of the hindgut. Myo31DF encodes an unconventional myosin I. In embryos homozygous for Myo31DF, LR asymmetry of the hindgut becomes mirror image of that of wild-type. In embryos homozygous for DE-Cad, the LR symmetry of the hindgut becomes randomized. We also found that Wnt and JNK signalling pathways need to be normally activated for the LR asymmetric development of the anterior part of the embryonic gut. The LR asymmetric morphogenesis of the anterior midgut depends on Myosin II activity. Besides these mutants, mapping and molecular characterizations of other mutants are in progress.
Among these mutation, Myosin31DF (Myo31DF) (also called Myosin ID) and Drosophila E-Cadherin (DE-Cad) affect the LR symmetry of the hindgut. Myo31DF encodes an unconventional myosin I. In embryos homozygous for Myo31DF, LR asymmetry of the hindgut becomes mirror image of that of wild-type. In embryos homozygous for DE-Cad, the LR symmetry of the hindgut becomes randomized. We also found that Wnt and JNK signalling pathways need to be normally activated for the LR asymmetric development of the anterior part of the embryonic gut. The LR asymmetric morphogenesis of the anterior midgut depends on Myosin II activity. Besides these mutants, mapping and molecular characterizations of other mutants are in progress.
Computer modeling of the LR asymmetric morphogenesis of the hindgut epithelium
To understand the contribution of PCC to the LR asymmetric morphogenesis, we introduced computer modeling approach. We simulated the PCC formation in a model epithelia tube in silico. Our model revealed that PCC was sufficient to induced LR asymmetric rotation of the hindgut epithelial tube, which is observed in vivo.
Mechanobiology of LR asymmetric development
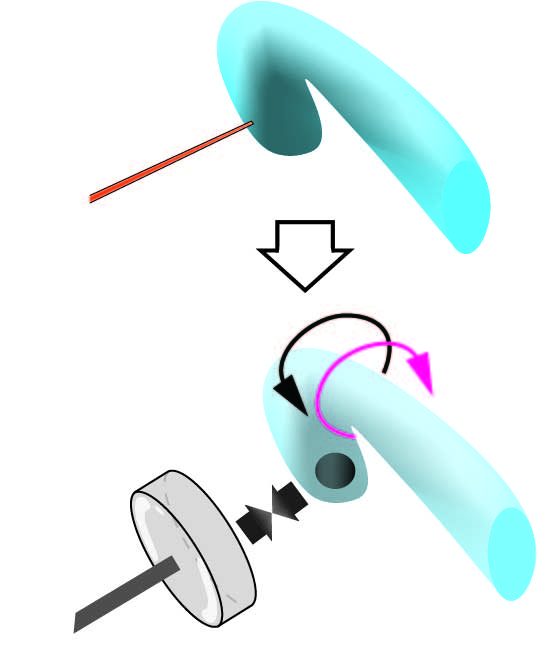
During the LR asymmetric morphogenesis, the hindgut epithelial tube makes a 90 degree counterclockwise rotation. To understand the mechanisms of this rotation at the cellular level, we thought that the mechanical force needs to be measured. We developed a novel method to measure the torque of the hindgut counterclockwise rotation, which applies the magnetic beads.
Before the rotation of the hindgut, the anterior end is curved towards the ventral side of the embryos, although it is LR symmetric. Magnetic beads were injected into the anterior end of the hindgut lumen. Using neodymium magnets, we induced counterbalancing magnetic force against the twisting of the hindgut. As a reference of how much force this counterbalancing created, we measure the magnetic force between the magnetic beads and the neodymium magnet in viscous fluid using Stokes’ low.
We successfully quantified the magnetic force required to stop the contortion of the hindgut, which is equivalent to the force generated by the hindgut for the twisting. These values are crucial for performing quantitative genetic analysis of mechanical force inducing the LR asymmetric morphogenesis. Currently, we are building a computer model reflecting the value of the mechanical force.
Notch signaling
Identification of novel Notch signal components by genetic screen of maternal genes
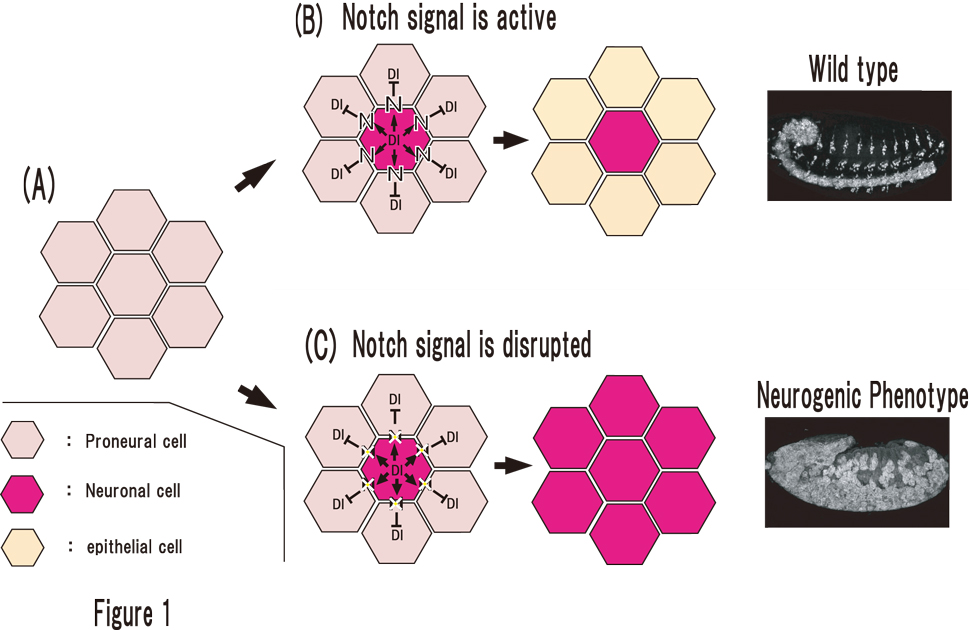
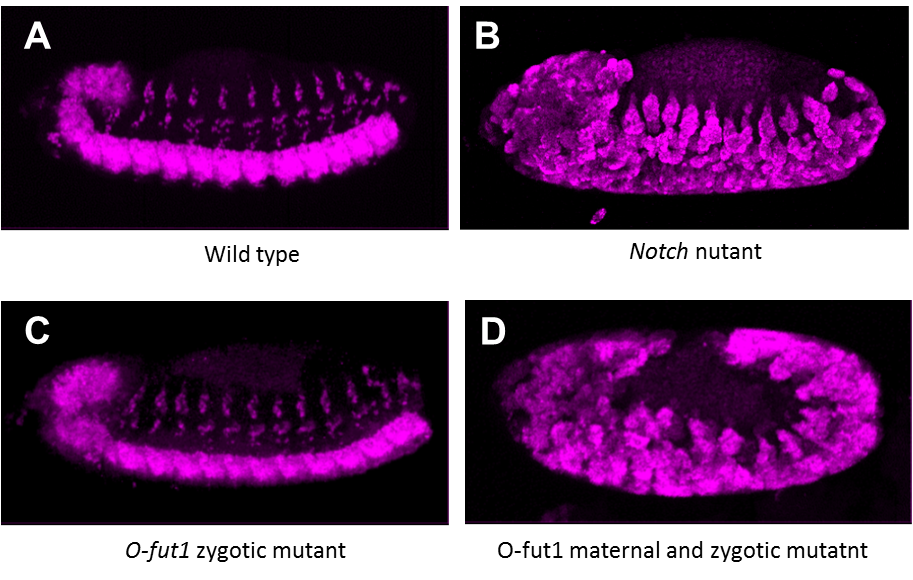
To identify novel components of Notch signaling, we conducted a genetic screen for mutants that show maternal neurogenic phenotype. We screened about 500 mutagenized second chromosomes (left-arm) and obtained novel mutations showing maternal neurogenic phenotype. The expression of Notch signal target genes were severely reduced in embryos homozygous for one of these mutants lacking its maternal contribution. Currently, we are mapping the genetic locus of this mutation.
Analysis of pecanex gene function in Notch signaling
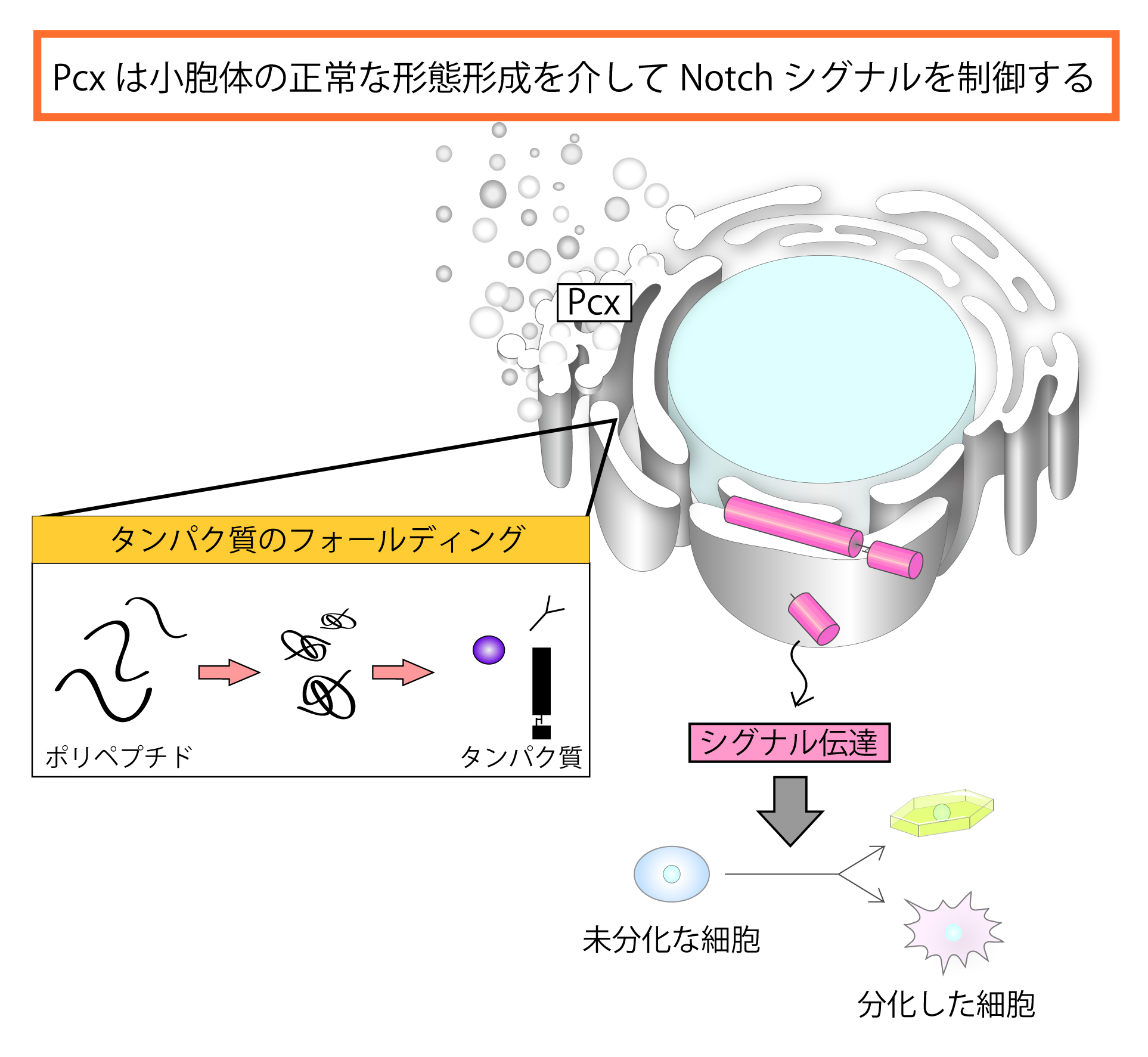
pecanex (pcx) is a maternal neurogenic gene Recently, we showed that the expression of Notch target genes, Enhancer of split m8 and single-minded is decreased in embryos homozygous for pcx lacking its maternal contribution.
From these data, we concluded that Pcx is a component of Notch signaling. We also found that the ER is enlarged in these embryos, and Pcx is an ER resident protein. In addition, our genetic analysis suggested that pcx functions upstream of an activated form of Notch (NICD) in Notch signal receiving cells. From these results, we speculated that Pcx may play a role in maturation of Notch in the ER.
Consistent with this idea, we found that overexpression of an activated form of XbpI or a dominant negative form of Hsc70, both of which activates UPR (unfolded protein response, one of the early cellular responses to ER stress) rescued the maternal neurogenic phenotype of pcx. Therefore, we propose that Pcx is involved in some ER function, which is required for the proper folding of Notch. To identify genes encoding components associated with the Pcx function, we are conducting a genetic modifier screen based on a pcx phenotype.
Consistent with this idea, we found that overexpression of an activated form of XbpI or a dominant negative form of Hsc70, both of which activates UPR (unfolded protein response, one of the early cellular responses to ER stress) rescued the maternal neurogenic phenotype of pcx. Therefore, we propose that Pcx is involved in some ER function, which is required for the proper folding of Notch. To identify genes encoding components associated with the Pcx function, we are conducting a genetic modifier screen based on a pcx phenotype.
Analysis of O-fucosyltransferase1 function in Notch signaling
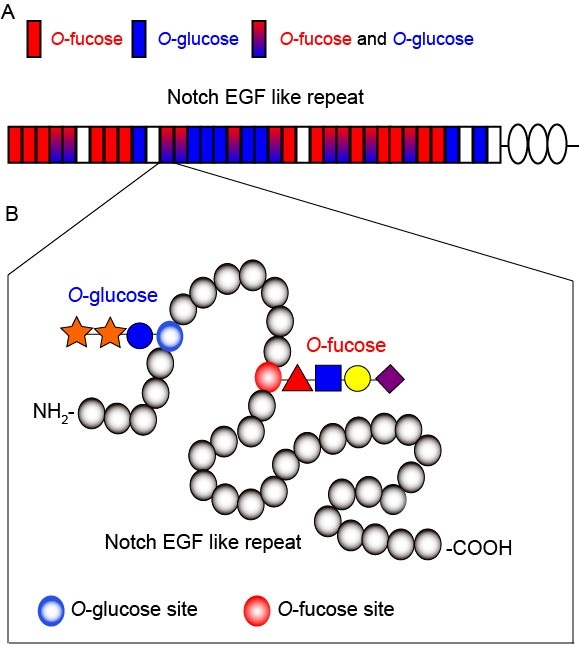
Previously, we identified O-fucosyltransferase1 (O-fut1) gene, which catalyzes O-fucose modification of epidermal growth factor (EGF)-like repeats with a consensus sequence in the extracellular domain of Notch. We are interested in the function of O-fut1 and O-fucosylation of Notch in Notch signaling.
The O-fucose modification by O-fut1 is required for the regulation of Notch-ligand interaction by fringe, which encodes an enzyme that adds GlcNAc to this O-fucose. However, the specific function of the monosaccharide O-fucose is controversial. We found that monosaccharide O-fucose is essential only at high temperature (29 oC) in fringe-independent Notch signaling, although it is dispensable at the normal temperature (25 oC). Thus, we speculated that the monosaccharide O-fucose may play a role in promoting the proper folding of Notch extracellular domain.
