Contents
Research Activities
Professor / Hisao MASUKATA Ph.D.
Research Projects: Mechanisms of eukaryotic DNA replication and genome integrity
Summary of Research Activity
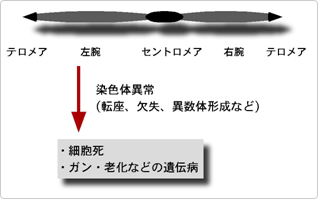
DNA replication is an essential reaction to maintain the genetic information. For complete duplication of large and complex genome in eukaryotic cells, DNA replication initiates at large number of sites, called replication origins, and the initiation process is tightly regulated by the cell cycle. We would like to understand the molecular mechanism of initiation of DNA replication, espetially regulation by cell cycle and chromatin sturctures. We are also interested in reactions tightly coordinated with DNA replication. The stalled replication fork need to be recovered by repair and recombination. Failure in the recovery results in cell death or genome instability that may cause a cancer in multicellular organisms. Sister chromatid cohesion has to be established during replication to separate the duplicated chromosomes. To understand these mechanisms at a molecular level, we use fission yeast, Schizosaccharomyces pombe, which is a suitable model organism to study the fundamental and general biological functions, and an in vitro system of Xenopus egg extracts (updated on 2012.8.23).
Project 1. Genome-wide identification of replication origins
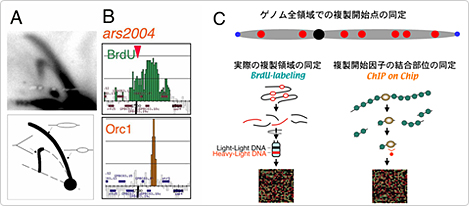
Chromosomes are confined into different chromatin structures relating with chromosome functions, such as centromere, mating type locus and telomeres. We are studying distribution of replication origins on fission yeast chromosomes by means of ChIP-chip analysis of replication factors and mapping of nascent DNA synthesized in early S phase (BrdU-chip) using high resolution tiling array (colaborating with Katsuhiko Shirahige). We identified 460 intergenic regions where Orc1 and Mcm6 are colocalized, as pre-replicative complex (pre-RC) sites. Among them 307 are active in early S phase (Early origins), while 153 do not fire in early S phase (called as Late origins). Interestingly, heterochromatin regions behave differently each other. Pre-RCs distribute randomly, while early and late origins tend to cluster each other, suggesting choromosome domains regulating initiation of replication. The heterochromatins at pericentromere and the mating-type locus replicate early S phase, while sub-telomeres replicate very late (Hayashi et al, EMBO J. 26, 1327-1339, 2007; Hayashi et al. Nat Cell Biol, 2009; Tazumi et al. Genes Dev, in press).
Project 2. Molecular mechanism of activation of replication origins
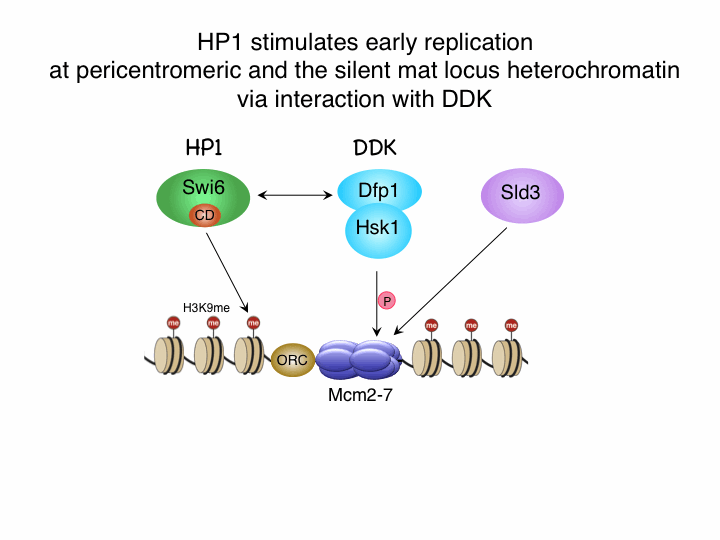
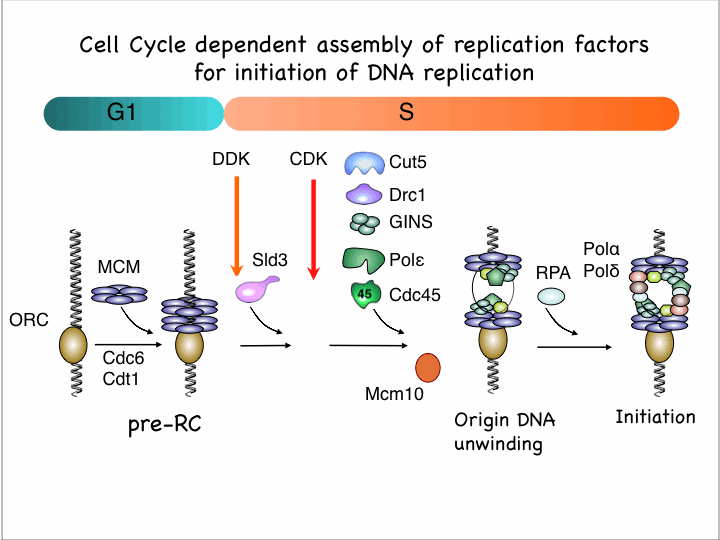
Replication origins are recognized by origin recognition complex (ORC) and minichromosome maintenance (MCM) complex is loaded depending on loading factors Cdc18/Cdc6 and Cdt1 in G1 phase. Assembly of several initiation factors is necessary for initiation of replication in S phase and this process is regulated by two conserved proteins kinases, cyclin-dependent kinase (CDK) and Dbf4-depedent kinase (DDK). By means of thermosensitive mutants of initiation factors, we found that factors assembly onto replication origins in distinct order; first Sld3 is loaded depending on DDK, then GINS, Cut5 are recruited depenmding on CDK, and depending on all these Cdc45 is loaded onto replication origin(Yabuuchi, et al. EMBO J. 25, 4663-4674, 2006).
Project 3. Mechanisms to maintain genome stability
(c)2006 Labolatory of Molecular Genetics